by: Estefania Gatton
A Song of LUCA and the Genetic Web
Before the first eye opened to the Ancient Sun, there was a whisper. Somewhere in the dark vents of a young ocean, a single spark took hold. We call this spark LUCA, the Last Universal Common Ancestor. It was a humble, single-celled existence. Yet, it carried within a unique feature: a language. A code as simple as itself, written in four letters, that would eventually describe every fingerprint, every leaf, and every human thought.
Every creature that has ever existed is a variation of that original theme. When you look at the jaguar moving through the shadows, the Ceiba camba tree—with the most beautiful flowers you can see—or even a microscopic yeast cell in a Biology 101 lab session, you are looking at a relative.
The thread of life is a live, woven web. Beyond the slow, steady march of traditional inheritance, there is a hidden pulse thrumming through the Earth. Genetics works as a world-wide system, an open-source library where the walls are made of mist.
Within the quiet architecture of a single cell, the code is never still. It rewrites history as it is being lived. Through the restless movement of transposable elements, life is reinventing itself from within. These “jumping genes” are the librarians of the deep code, moving verses from one chapter to another, ensuring the story never grows stagnant.
This code does not recognize the boundaries we draw. A bacterium whispers a secret of resilience to a fungus; a viral sequence leaves a gift of ancient immunity within a mammal. Across populations and even across different species, the original song from LUCA still plays in an interconnected biomachine, sharing an ancient memory through the currents of Horizontal Gene Transfer.
Powerful secrets are being revealed; others remain hidden in the folds of our DNA. Every creature is a vessel for these wandering verses, carrying the potential to adapt, to heal, and to thrive in worlds we have yet to imagine.
We are the latest verse in an epic that began in the dark solitude of the sea, written in the ink of LUCA, and carried forward by the restless wind of spreading genes, a story refusing to stay on the page of a book.
“Jumping genes”: Modifying the Future of Evolution and Life
For over half a century, the central dogma of biology was taught as a rigid, linear progression. DNA was the master blueprint, such as a stable, ancient library of instructions passed from generation
to generation with the sacred duty of remaining unchanged, like book pages being duplicated by a copy machine. In this traditional view, mutations were accidents, “typos” in the code that were sometimes detrimental. However, as our sequencing technology has reached a resolution once thought impossible, we have discovered a chaotic truth: our DNA is not a static document. It is a shifting, breathing mosaic.
Nearly 50% of the human genome is composed of Transposable Elements (TEs) (1). TEs are sequences of DNA that can move from one location to another (2). Long dismissed by 20th-century scientists as “junk DNA” or “genomic parasites,” these molecular acrobats are now recognized as the engineers of life (3). They are the primary drivers of evolution in the wild and, increasingly, the foundation of the most sophisticated tools in modern biotechnology.
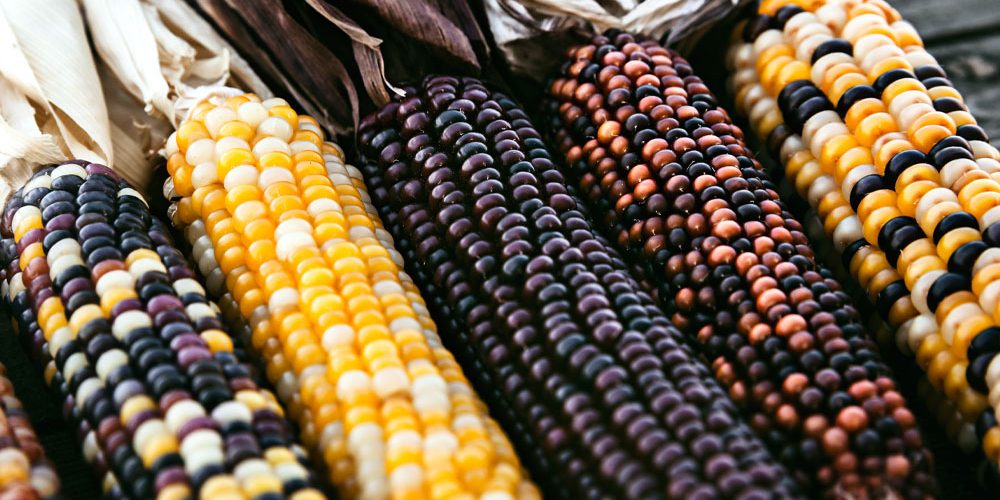
The Maverick of Maize
The story of transposable elements began not in a high-tech lab, but in a 1940s cornfield. Geneticist Barbara McClintock noticed that the color patterns in maize kernels didn’t follow the predictable laws of Mendelian inheritance. She realized that certain genetic segments were “jumping” into color-producing genes, switching them off or on depending on where they landed (4). McClintock’s “jumping genes” were initially met with deep skepticism; the idea of a fluid genome threatened the very stability of biological science. Yet, she was eventually awarded the Nobel Prize, proving that the genome is a dynamic organ capable of reorganizing itself in response to environmental stress.
To understand how TEs function, we must distinguish between their two primary methods of travel. Class I Retrotransposons (Copy-Paste): They transcribe themselves into RNA, which is then converted back into DNA and inserted into a new location. And Class II DNA Transposons (Cut Paste): Using an enzyme called transposase, they physically excise themselves from one spot and move to another (2).
Real-World Evolution
In nature, this scrambling of genes means survival. The Peppered Moth is a classic example from biology courses, and now it has a twist. During the Industrial Revolution, these moths rapidly evolved dark wings to camouflage against soot-covered trees. Recent genomic analysis revealed this wasn’t a simple point mutation; it was a transposable element jumping into a gene called cortex, which regulates wing patterning (5). Similarly, the deep crimson of a Blood Orange is triggered by a retrotransposon that sits near a pigment gene. When the fruit is stressed by cold weather, the “jumping gene” activates, turning on the production of anthocyanins (6). TEs are built in mechanisms for rapid adaptation.
The Mosaic Brain
For decades, the “one genome, one body” rule was a pillar of biology. We assumed the DNA in your heart was identical to the DNA in your brain. However, landmark research proved otherwise. By using fluorescent markers, scientists saw LINE-1 (L1) retrotransposons jumping into the DNA of developing neurons (7). These “jumps” mean that every neuron in your head could have a slightly different genetic code. But are they just causing random chaos? In 2025, researchers at the forefront of cerebral organoid technology (mini-brains grown in labs) found interesting results. They used a tool called CRISPRi to “silence” these jumping genes to see what would happen. The results were shocking: Without the jumping genes, the organoids couldn’t grow to their full size. The “blueprint” for a human-sized brain simply didn’t work without the movement of these elements (8).
The “Sleeping Beauty” System
One of the most poetic success stories in biotech is the Sleeping Beauty (SB) transposon. Millions of years ago, this DNA transposon became “extinct” in the genomes of fish, rendered inactive by mutations. In the late 1990s, scientists reawakened it by reconstructing its ancestral sequence (9). Today, SB is a leading non-viral vector for gene therapy. In early 2026, new data from pediatric clinical trials showed that Sleeping Beauty-based CAR-T cell therapy was achieving remission rates in leukemia patients at a fraction of the cost of traditional viral-based therapies (10). Because SB is a simple plasmid-based system, it avoids the massive manufacturing hurdles and immune reactions associated with using modified viruses like HIV or Adenovirus to deliver genes.
The “PiggyBac” Advantage
Named for its ability to carry massive genetic payloads, the PiggyBac system is prized for its “scarless” engineering. Most gene-editing tools leave behind a small “scar” of extra DNA when they move. PiggyBac, however, can insert a gene and later be removed so cleanly that the original sequence is restored perfectly. This is currently the gold standard for creating induced Pluripotent Stem Cells (iPSCs), allowing scientists to turn a patient’s skin cells into heart or nerve cells without leaving any permanent genetic debris behind (11).
These examples make us think: What other treasures can we find in the future between Transposable Elements?
De-extinction and Rewilding the Future
Transposable elements are also the silent engines behind the most ambitious ecological project of the century: De-extinction. Companies like Colossal Biosciences are not just trying to simply
clone a Woolly Mammoth; they are trying to engineer an elephant that can survive in the Arctic (12). To do this, they use TEs as high-speed delivery vehicles. By using multiplexed transposon systems, they can integrate hundreds of cold-adaptation genes, regulating different structures and mechanisms from hair length to subcutaneous fat, into the Asian Elephant genome simultaneously.
This isn’t just a science experiment; the real purpose is an attempt to restore the “mammoth steppe” ecosystem, which could sequester billions of tons of carbon and help stabilize the melting permafrost.
Aging and “Inflammaging” Double-Edged Sword
However, there is a dark side to this genomic fluidity of the TEs. We can think as Life and Evolution as constant battle between the need for adaptation and the need for stability. As we age, our cellular “police”, the epigenetic markers that keep TEs locked in place, begin to fail. When LINE-1 elements start jumping uncontrollably in an aging body, the consequences are dire. The cell perceives the stray DNA and RNA produced by these genes as a viral infection. This triggers a massive, chronic inflammatory response known as “inflammaging.” This state of permanent high alert is now believed to be a primary driver of Alzheimer’s, Parkinson’s, and Type 2 Diabetes (13).
In the other side, as of March 2026, a groundbreaking $22 million clinical trial funded by ARPA-H is testing a radical hypothesis. Researchers are testing a drug called TPN-101, that is an antiretroviral originally designed to treat HIV. The goal is to “freeze” the jumping genes by blocking the enzymes they use to move. If successful, we may be able to treat the one of the root causes of aging itself, silencing genomic chaos and extending the human health span by decades (14).
Horizontal Gene Transfer
While TEs rearrange the code within a single lineage, Horizontal Gene Transfer (HGT) takes this fluidity to a global scale. HGT is the process by which genetic material moves between different species (15). A phenomenon once thought to be exclusive to bacteria but now recognized as a major force in the evolution of complex animals and plants. Research has shown that this “genetic theft” allows for near-instantaneous adaptation. For example, the Bov-B retrotransposon has been found to have jumped between snakes and cows, likely carried by ticks (16). In other cases, plants have “borrowed” genes from bacteria to survive in toxic soils (17). By sharing pieces of DNA across the borders of species, life ensures that a successful survival strategy developed by one organism can eventually benefit the entire biosphere.
Conclusions
We are currently standing at the end of the Static age of genetics. We now understand that a genome that cannot change is a genome that cannot survive. Transposable elements and HGT
remind us that the code of life is a constant, shifting dialogue with the present environment. Whether we are reawakening ancient fish DNA to cure cancer, using “jumping” logic to bring back the Woolly Mammoth, or taking HIV drugs to stop the clock on aging, we are finally learning to dance with the chaos of our own DNA. We are no longer just the product of our genes; we are becoming their editors, their directors, and their future.
photo
References:
1. Ayarpadikannan, S., & Kim, H. S. (2014). The impact of transposable elements in genome evolution and genetic instability and their implications in various diseases. Genomics & informatics, 12(3), 98–104. https://doi.org/10.5808/GI.2014.12.3.98
2. Wells, J. N., & Feschotte, C. (2020). A Field Guide to Eukaryotic Transposable Elements. Annual review of genetics, 54, 539–561. https://doi.org/10.1146/annurev-genet-040620- 022145
3. Senft, A.D., Macfarlan, T.S. Transposable elements shape the evolution of mammalian development. Nat Rev Genet 22, 691–711 (2021). https://doi.org/10.1038/s41576-021- 00385-1
4. McClintock, B. (1950). The origin and behavior of mutable loci in maize. 5. Hof, A., Campagne, P., Rigden, D. et al. The industrial melanism mutation in British peppered moths is a transposable element. Nature 534, 102–105 (2016). https://doi.org/10.1038/nature17951
6. Butelli, E., Licciardello, C., Zhang, Y., Liu, J., Mackay, S., Bailey, P., Reforgiato-Recupero, G., & Martin, C. (2012). Retrotransposons control fruit-specific, cold-dependent accumulation of anthocyanins in blood oranges. The Plant cell, 24(3), 1242–1255. https://doi.org/10.1105/tpc.111.095232
7. Muotri, A. R., Chu, V. T., Marchetto, M. C., Deng, W., Moran, J. V., & Gage, F. H. (2005). Somatic mosaicism in neuronal precursor cells mediated by L1 retrotransposition. Nature, 435(7044), 903–910. https://doi.org/10.1038/nature03663
8. Adami, A., Garza, R., Gerdes, P., Johansson, P. A., Dorazehi, F., Koutounidou, S., Castilla Vallmanya, L., Atacho, D. A. M., Sharma, Y., Johansson, J. G., Tam, O., Kirkeby, A., Barker, R. A., Hammell, M. G., Douse, C. H., & Jakobsson, J. (2025). LINE-1 retrotransposons mediate cis-acting transcriptional control in human pluripotent stem cells and regulate early brain development. Cell genomics, 5(10), 100979.
9. Ivics, Z., Hackett, P. B., Plasterk, R. H., & Izsvák, Z. (1997). Molecular reconstruction of Sleeping Beauty, a Tc1-like transposon from fish, and its transposition in human cells. Cell, 91(4), 501–510. https://doi.org/10.1016/s0092-8674(00)80436-5
10. Magnani, C. F., Gaipa, G., Lussana, F., Belotti, D., Gritti, G., Napolitano, S., Matera, G., Cabiati, B., Buracchi, C., Borleri, G., Fazio, G., Zaninelli, S., Tettamanti, S., Cesana, S.,
Colombo, V., Quaroni, M., Cazzaniga, G., Rovelli, A., Biagi, E., Galimberti, S., … Biondi, A. (2020). Sleeping Beauty-engineered CAR T cells achieve antileukemic activity without severe toxicities. The Journal of clinical investigation, 130(11), 6021–6033. https://doi.org/10.1172/JCI138473
11. Zhao, S., Jiang, E., Chen, S., Gu, Y., Shangguan, A. J., Lv, T., Luo, L., & Yu, Z. (2016). PiggyBac transposon vectors: the tools of the human gene encoding. Translational lung cancer research, 5(1), 120–125. https://doi.org/10.3978/j.issn.2218-6751.2016.01.05
12. Woolly Mammoth De-extinction Project & Process https://colossal.com/mammoth/ 13. Gorbunova, V., Seluanov, A., Mita, P., McKerrow, W., Fenyö, D., Boeke, J. D., Linker, S. B., Gage, F. H., Kreiling, J. A., Petrashen, A. P., Woodham, T. A., Taylor, J. R., Helfand, S. L., & Sedivy, J. M. (2021). The role of retrotransposable elements in ageing and age-associated diseases. Nature, 596(7870), 43–53. https://doi.org/10.1038/s41586-021-03542-y 14. 2026. ARPA-H awards research teams to add more healthy years to Americans’ lives as they age https://arpa-h.gov/news-and-events/research-teams-add-more-healthy-years americans-lives-they-age
15. Burmeister A. R. (2015). Horizontal Gene Transfer. Evolution, medicine, and public health, 2015(1), 193–194. https://doi.org/10.1093/emph/eov018
16. Puinongpo, W., Singchat, W., Petpradub, S., Kraichak, E., Nunome, M., Laopichienpong, N., Thongchum, R., Intarasorn, T., Sillapaprayoon, S., Indananda, C., Muangmai, N., Suntrarachun, S., Baicharoen, S., Chanhome, L., Peyachoknagul, S., & Srikulnath, K.
(2020). Existence of Bov-B LINE Retrotransposons in Snake Lineages Reveals Recent Multiple Horizontal Gene Transfers with Copy Number Variation. Genes, 11(11), 1241. https://doi.org/10.3390/genes11111241 17. Quispe-Huamanquispe, D. G., Gheysen, G., & Kreuze, J. F. (2017). Horizontal Gene Transfer Contributes to Plant Evolution: The Case of Agrobacterium T-DNAs. Frontiers in plant science, 8, 2015. https://doi.org/10.3389/fpls.2017.02015